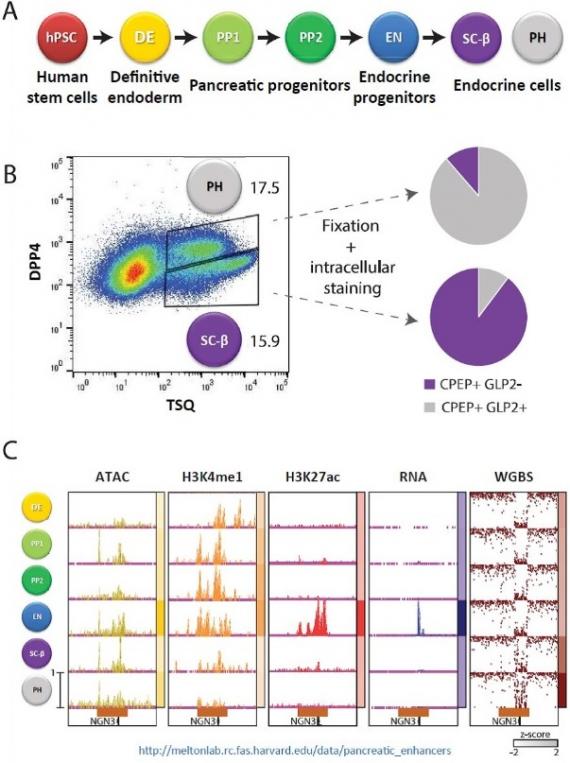
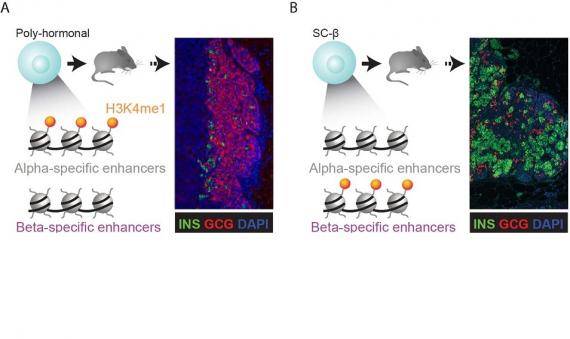
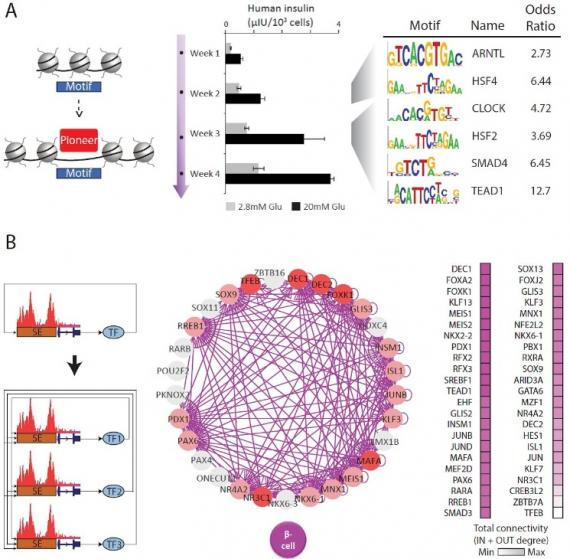
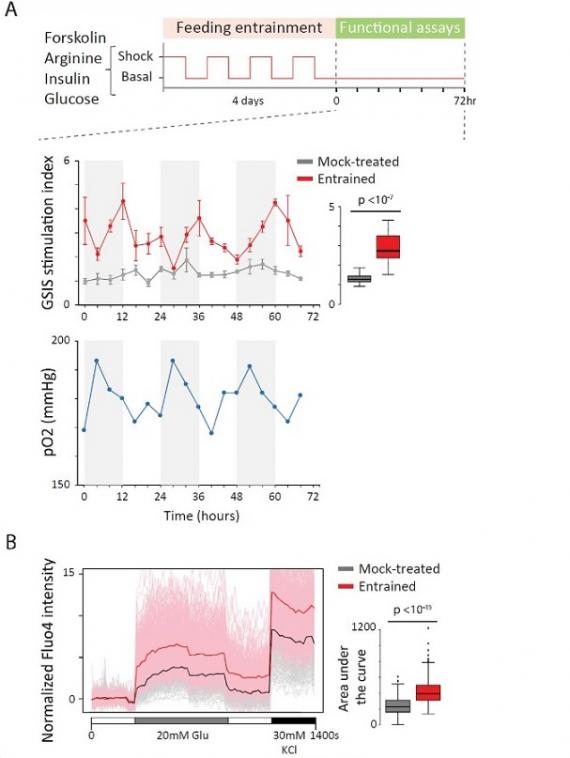
Organoid Maturation by Circadian Entrainment
Juan R. Alvarez-Dominguez,*1 Julie Donaghey,1 Niloofar Rasouli,1 Jennifer H. R. Kenty,1 Aharon Helman,1 Jocelyn Charlton,1,2 Juerg R. Straubhaar,1 Alexander Meissner,1,2 Douglas A. Melton,1
1 Department of Stem Cell and Regenerative Biology, Harvard Stem Cell Institute, Harvard University, Cambridge, MA, USA
2 Department of Genome Regulation, Max Planck Institute for Molecular Genetics, Berlin, Germany
* Corresponding author: juanralvarez@fas.harvard.edu
Submitted: Aug 4, 2020; Published online: Oct 24, 2020
Abstract
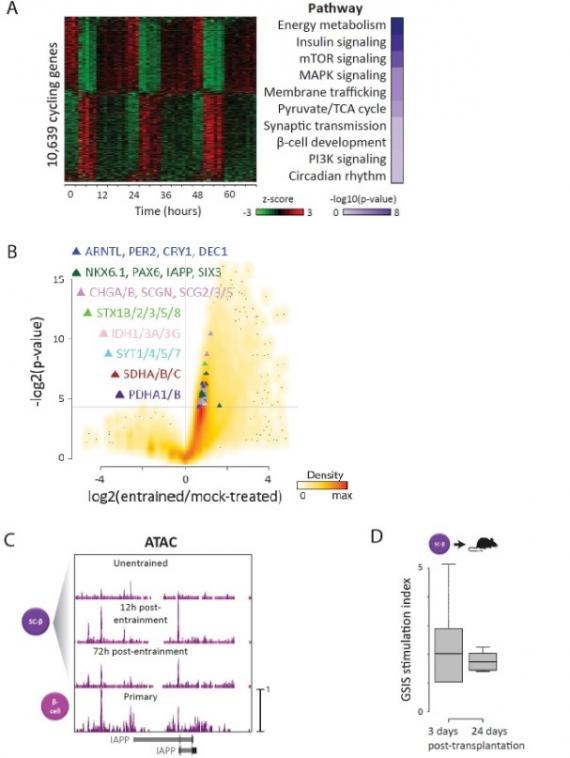
Stem cell-derived tissues that recap endogenous physiology are key for regenerative medicine. Yet, most methods yield products that function like fetal, not adult tissues. Organoids are typically grown in constant environments, while our tissues mature along with behavioral cycles. Here, we show that inducing circadian rhythms in pancreatic islet organoids, by entraining them to daily feeding-fasting cycles, elicits their metabolic maturation. Our results show that rhythms can be harnessed to further functional maturation of organoids destined for human therapeutics.