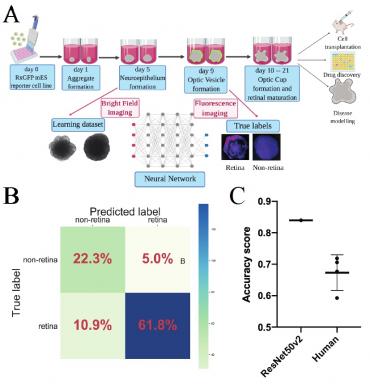
Deep Learning-Based Analysis of the Differentiation in 3D Retinal Organoids
Evgenii Kegeles,*1,2 Anton Naumov,3 Tatiana Perepelkina,1 Julia Oswald,1 Evgeniy Karpulevich,2,3,4 Pavel Volchkov,2 and Petr Baranov1
1 Schepens Eye Research Institute of Massachusetts Eye and Ear, Harvard Medical School Affiliate, Boston, MA, USA
2 Moscow Institute of Physics and Technology, Moscow, Russia
3 Ivannikov Institute for System Programming of the Russian Academy of Sciences, Moscow, Russia
4 National Research Center “Kurchatov Institute”, Moscow, Russia
* Presenting and corresponding author: kegeles.ea@phystech.edu
Submitted: Jun 11, 2020; Published online: Jul 27, 2020